Log-linear Graphical Model
: inferring probabilistic conditional independency from combinatorial regulation of transcription factors
To run LLGM,
please log in OpenLooper and visit this page again.
How to run LLGM pic.twitter.com/ll7hy4ZbDb
— OpenLooper (@w3olp) May 19, 2020
Introduction
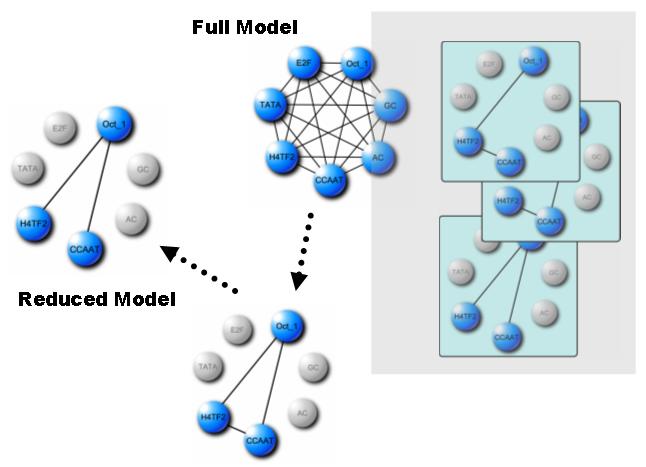 Transcription factors (TFs) regulate gene expressions by intricate combinatorial interactions. Computational modeling to infer the interactions is essential to understand the nature of transcriptional regulation. Log-linear Graphical Model (LLGM) deals with spurious TF interactions that may lead to deceptive inferences. Given a discrete (0 or 1) input data matrix of TF-DNA binding instances, the LLGM estimates the probabilistic conditional independency for detecting the spuriousness, which is known as "Simpson's Paradox".
Transcription factors (TFs) regulate gene expressions by intricate combinatorial interactions. Computational modeling to infer the interactions is essential to understand the nature of transcriptional regulation. Log-linear Graphical Model (LLGM) deals with spurious TF interactions that may lead to deceptive inferences. Given a discrete (0 or 1) input data matrix of TF-DNA binding instances, the LLGM estimates the probabilistic conditional independency for detecting the spuriousness, which is known as "Simpson's Paradox".
Instruction
The format of the input matrix file:- "0 1" (=presence) of TF1
- "1 0" (=absence) of TF2
- "1 0" (=absence) of TF3.
Two thresholds are required for running:
- p-value cutoff for the test of deviance of a current RM (reduced model) from the FM (full model)
- p-value cutoff for the test of deviance of a current RM (reduced model) from the previous RM
NOTE:
Reference
1. Lauritzen, S.L., "Graphical Models", Oxford University Press, 1996
2. Christensen, R., "Log-Linear Models and Logistic Regression", Springer-Verlag, 1997
3. Park SJ, Umemoto T, Saito-Adachi M, Shiratsuchi Y, Yamato M, Nakai K, "Computational Promoter Modeling Identifies the Modes of Transcriptional Regulation in Hematopoietic Stem Cells", PLoS ONE 9(4): e93853, 2014